Antibiotics Resistance Profile of Staphylococci Isolated from Urogenital Infections and Toxins Production of Staphylococcus aureus Strains
2 Polytechnic School of Abomey-Calavi, University of Abomey-Calavi, Benin
3 Laboratoire de Microbiologie et des Technologies Alimentaires, Faculté des Sciences et Techniques (FAST), Université d’Abomey-Calavi, 01 BP: 2009 Cotonou, Benin
4 Laboratoire de Recherche en Biologie Appliquée (LARBA), Ecole Polytechnique d’Abomey-Calavi (EPAC), Université d’Abomey-Calavi, 01 BP. 2009 Cotonou, Benin
Citation: Sina H, et al. Antibiotics Resistance Profile of Staphylococci Isolated from Urogenital Infections and Toxins Production of Staphylococcus aureus Strains. Ann Med Health Sci Res. 2018; 8: 29-34
This open-access article is distributed under the terms of the Creative Commons Attribution Non-Commercial License (CC BY-NC) (http://creativecommons.org/licenses/by-nc/4.0/), which permits reuse, distribution and reproduction of the article, provided that the original work is properly cited and the reuse is restricted to noncommercial purposes. For commercial reuse, contact reprints@pulsus.com
Abstract
Aim: The aim of this study was to investigate the antibiotic resistance of staphylococci and seek toxin production by Staphylococcus aureus strains isolated from urogenital infections. Material and Methods: The staphylococci strains were isolated from urogenital samples collected from hospitalized patients or not. The antibiotic susceptibility was performed by the diffusion method and the search of production of toxin by S. aureus was done by radial immunoprecipitation technique. Results: Out of the 1904 samples analyzed, 80 staphylococci strains were isolated. The major (70%) part of the positive samples were coagulase-negative staphylococci composed of S. saprophyticus (50.0%), S. epidermidis (16.25%), S. xylosus (2.5%), and S. haemolyticus (1.25%). S. aureus was the unique coagulase positive strains. It was observed a multi-resistance of the isolated strains to beta-lactams, aminoglycosides, tetracycline, and co-trimazole. All the S. haemolyticus and S. xylosus strains were resistant to methicillin. Nitrofurantoin was the most active molecule in all kind of strains. There was no methicillin-resistant S. aureus producing Panton-Valentine Leukocidin (PVL) detected but all the S. aureus producing PVL were community methicillin-sensitive S. aureus. Most of the tested strains produced ETB (83.33%) and ETA (45.33%). Conclusion: The presence of multidrug resistance staphylococci strains producing toxins indicate an existence of potential reservoir of virulent antibiotics resistance stains in the community.
Keywords
Urogenital infections; Antibiotic resistance; Staphylococcus; PVL; Exfoliatin; MRSA; Benin
Introduction
Bacteria are present on our skin, in our mucous membranes and although the vast majority of them are beneficial to us, there are also pathogens. [1,2] To control these pathogenic bacteria, antimicrobial agents are often used since 1940 without clearing those bacteria. Indeed, bacteria develop several strategies (natural and/or acquired) to overcome antibiotics effects. [1] However, the acquired resistance is the most spread in bacterial and has now become a global phenomenon. [3]
The acquired mechanisms of resistance to antibiotics include the antibiotic target modification mechanism especially observed in Gram-positive cocci. [4,5] Among these target modification mechanisms, the production of a new penicillin-binding protein (PBP) such as PBP2a or PBP2’ with little affinity for betalactams and leading to resistance to methicillin is distinguished. [6,7] The emergence of resistant staphylococci strains to betalactams by this mechanism is therefore of concern since the beta-lactam antibiotics are considered very efficacy with few side effects. [1,8]
Apart from antibiotic resistance, staphylococci, especially S.aureus, also have a high ability to produce virulence factors such as Panton and Valentine Leucocidin (PVL), exfoliatins (ETA and ETB), and so on. PVL is a two-compound toxin (synergistic action of two LukS-PV and LukF-PV compounds) belonging to the family of pore-forming toxins whereas exfoliatins are epidermolytic toxins. [9] The prevalence of staphylococcal strains producing PVL is constantly increasing worldwide. [10] More worryingly, we have the worldwide emergence of toxinogenic and methicillin-resistant strains of S. aureus. [11]
staphylococci including coagulase positive (S. aureus) and Coagulase negative (S. saprophyticus, S. epidermidis, etc.) [12] are the second class of bacteria isolated from urogenital infections and particularly urinary tract infections, occupy an important place in the nosocomial and community acquired infections. [13] Indeed, urinary tract infections are a major public health problem and the consequences of untreated or poorly treated urogenital infections causing cystitis, abortion, sterility and even death. [14]
In West Africa, staphylococcal infections are widespread and are often associated with urogenital infections [15] In Benin, the place of urogenital infections is no less important because apart from Sexually Transmitted Diseases, they represent 2.1% of the main diseases. Studies have been carried out on the antimicrobial resistance of S. aureus strains isolated from urinary and/or genital infections and their toxins production. Unfortunately, to date very few studies have taken into account both S. aureus and Coagulase negative Staphylococcus. [16] In order to fill these gaps and to monitor the evolution of bacterial resistance, this study was initiated to study the antibiotic resistance of staphylococci isolated from urogenital infections and investigate the production of toxins by S. aureus.
Material and Methods
Sample collection
Samples of 1904 hospitalized or untreated patients were collected at the National University Hospital of Cotonou (Benin) for four month (from November 2012 to March 2013) various bacteriological screenings in routine. Those samples were composed of urine (1036), sperm (292), urethral (244) and vaginal (332) samples. All the samples were collected following the standard recommendations of each sample. [17,18] Once collected, the samples were stored at 4°C until microbial analysis were performed for the research of Staphylococcus strains.
Staphylococcus strains identification
Standard microbiological methods for identification of microorganisms were applied [19] Briefly, all swabs were inoculated onto mannitol salt agar, incubated at 37°C and inspected. Any suspected colony was subcultured on Mueller- Hinton agar (bioMérieux, Marcy l’Etoile, France) and identified by subsequent Gram staining, catalase test, DNase test, susceptibility to novobiocin test and Slidex Staph Plus (bioMérieux, Marcy l’Etoile, France) and the coagulase test with the rabbit plasma [20] Finally, bacterial identification was completed by API Staph (bioMérieux, Marcy l’Etoile, France).
Antibiotics susceptibility
The susceptibility of the identified Staphylococcus strains to 19 antibiotics molecules was determined by the disc diffusion method of Kirby-Bauer on agar Mueller-Hinton (bioMérieux, Marcy l’Etoile, France) as recommended by the Antibiogram Committee of the French Microbiology Society. [21] After incubation time (18-24 h) at 37°C, inhibition zone was measured using graduated slide ruler applied in contact with the rear face of the petri dish. For susceptibility to oxacillin, inoculum of 107 CFU/ml was prepared and the plate was incubated at 37°C for 24 h on Mueller-Hinton agar + 3% NaCl.
The 19 tested antibiotics provided by Biomérieux (Marcy l’Etoile, France) were: Amoxicillin (AMX 30 µg), Oxacillin (OXA 5 µg), Penicillin G (PENI G 6 µg), Amoxicillin + clavulanic acid (AMC 20/10 µg), Cefoxitin (FOX 30 µg), Gentamicin (GM 10 UI), Tobramycin (TM 10 µg), Kanamycin (K 30 µg), Netilmycin (NET 30 µg), Erythromycin (ERY 5 µg), Spiramycin (SP 100 µg), Pristinamycin (PT 15 µg), Lincomycin (LIN 10 µg), Ciprofloxacin (CIP 5 µg), Nitrofurantoin (FT 300 µg), Cotrimazol (COT 25 µg), Chloramphenicol (C 30 µg), Tetracycline (TE 30 µg), and Doxycycline (DO 30 UI).
Toxin production
The capability of the isolated S. aureus strains to produce Panton-Valentine Leukocidin (PVL) and epidermolysins A (ETA) and B (ETB) was performed phenotypically by radial gel immuno-diffusion. Thus, those toxins (PVL, ETA and ETB) were evidenced from culture supernatants after 18-24 h of growth in Yeast Casamino-acid Pyruvate broth medium [22] by radial gel immuno-diffusion in 0.6% (wt/vol) agarose with component-specific rabbit polyclonal and affinity-purified antibodies. [23,24]
Statistical analysis
Microsoft Office Excel 2007 spreadsheet was used for statistical data processing. The SPSS (V 16 2007) and Graph Pad Prism 5 software was used to determine whether there is a relationship between the characters being compared. Thus, for comparison tests of positive isolates of each patient group, we used the Student T test, and the Fischer’s test for lower number series. P< 0.05 was considered statistically significant.
Results
Samples contamination
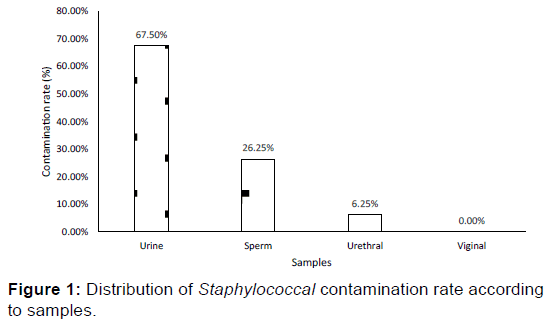
A total of 80 Staphylococcus strains were isolated mostly (95%) from non-hospitalized (community) patients sample. Those strains were isolated from urine (67.5%), sperm (26.25%), and urethral samples (6.25%). The vaginal samples were free of Staphylococcus strains [Figure 1]. The urine samples containing the Staphylococcus strains were at 61% collected from females.
Five different staphylococcal species were isolated at different rate [Table 1] and depend on the samples (P< 0.001). Among the isolated staphylococcal strains, only S. aureus (50.0%) and S. saprophyticus (50.0%) were hospital acquired whereas in the community, all five species were represented. The distribution of Staphylococcus species according to the origin of the samples was random [Table 2].
| Samples Staphylococcal strains |
Urethral % (n) |
Sperm % (n) |
Urine % (n) |
Total % (n) |
|---|---|---|---|---|
| S. aureus | 80% (04) | 4.76% (1) | 35.19% (19) | 30% (24) |
| S. epidermidis | 20% (01) | 23.81% (7) | 12.96% (7) | 16.25% (13) |
| S. saprophyticus | 00% (00) | 61.91% (13) | 50.0% (27) | 50% (40) |
| S. haemolyticus | 00% (00) | 00% (00) | 1.85% (1) | 1.25% (1) |
| S. xylosus | 00% (00) | 9.52% (2) | 00% (00) | 2.5% (2) |
| Total | 6.25% (5) | 26.25% (21) | 67.5% (54) | 100% (80) |
| p<0.001 | ||||
Table 1: Distribution of Staphylococcus strains according to species and collection site
| Variables | S. aur | S. epi | S. sap | S. hae | S. xyl | Total |
|---|---|---|---|---|---|---|
| Hospital acquired | 50.0% (02) | 00% (00) | 50.0% (02) | 00% (00) | 00% (00) | 05.0% (04) |
| Community acquired | 28.95% (22) | 17.11% (13) | 50.0% (38) | 01.31% (01) | 02.63% (02) | 95.0% (76) |
| Total | 30.0% (24) | 16.25% (13) | 50.0% (40) | 01.25% (01) | 02.5% (02) | 100% (80) |
Table 2: Distribution of Staphylococci strains according to the species and sample origin.
Susceptibility to antibiotics
Penicillin G was inactive on all Staphylococcus species isolated with a 90% resistance rate for S. saprophyticus and 100% for the other four species. The proportion of resistance to methicillin was 54.17% (S. aureus), 52.50% (S. saprophyticus), 38.46% (S. epidermidis) and 100% (S. haemolyticus, S. xylosus). Amoxicillin and amoxicillin + clavulanic acid were also inactive on the five Staphylococcus species but with lower resistance levels for amoxicillin + clavulanic acid.
All the aminoglycoside (gentamicin, kanamycin, tobramycin, and netilmicin) were globally inactive on the five Staphylococcus species with at least more than half of the strains of each species resistant to these molecules. S. epidermidis, S. haemolyticus, and S. xylosus were more resistant to aminoglycosides with KTG-phenotypic than S. saprophyticus (78.43%).
Macrolides and especially Lincosamides and Streptogramines were more active on the other four species. The lowest resistance levels for pristinamycin were recorded with S. xylosus (00%) and S. epidermidis (7.7%). Ciprofloxacin was inactive on the five Staphylococcus species with the lowest (47.50%) rate of resistance observed on S. saprophyticus and the highest (100%) with S. xylosus. Chloramphenicol maintained good activity on S. haemolyticus and S. xylosus (100%) and S. epidermidis (15.39% resistance) and was less active (>40%) on S. aureus and S. saprophyticus. All staphylococci strains were resistant to tetracycline and doxycycline and the most active molecule on all five Staphylococcus species was nitrofurantoin [Table 3]. Thus, the resistance levels of the five Staphylococcus species depended on the types of antibiotics used (P<0.001).
| Antibiotics | S. aureus (n=24) | S. saprophyticus (n=40) | S. epidermidis (n=13) | S. haemolyticus (n=1) | S. xylosus (n=2) |
|---|---|---|---|---|---|
| Penicillin G | 100% | 90.00% | 100% | 100% | 100% |
| Oxacillin | 58.33% | 57.50% | 38.46% | 100% | 100% |
| Céfoxitine | 54.17% | 52.50% | 38.46% | 100% | 100% |
| Amoxicillin | 83.33% | 70.00% | 92.31% | 100% | 100% |
| Amoxicillin + clavulanic acid | 58.33% | 52.50% | 53.85% | 100% | 100% |
| Gentamycin | 54.17% | 55.00% | 76.9% | 100% | 100% |
| Kanamycin | 75.00% | 55.00% | 84.61% | 100% | 100% |
| Tobramycin | 66.67% | 55,00% | 76.92% | 100% | 100% |
| Netilmicin | 54.17% | 55.00% | 69.23% | 100% | 100% |
| Erythromycin | 50.00% | 60.00% | 15.39% | 100% | 00% |
| Spiramycin | 45.8% | 3500% | 30.77% | 100% | 00% |
| Lincomycin | 25.00% | 30.00% | 15.39% | 100% | 50.00% |
| Pristinamycin | 20.83% | 22.5% | 7.7% | 100% | 00% |
| Ciprofloxacin | 54.17% | 47.50% | 61.54% | 100% | 50.00% |
| Chloramphenicol | 50.00% | 40.00% | 15.39% | 00% | 00% |
| Tetracycline | 83.33% | 82.50% | 100% | 100% | 100% |
| Doxycycline | 87.50% | 77.50% | 100% | 100% | 100% |
| Cotrimazole | 50.00% | 85.00% | 92.31% | 100% | 100% |
| Nitrofurantoin | 4.17% | 00% | 00% | 00% | 00% |
Table 3: Resistance profile of Staphylococcal strains isolated from urogenital infections.
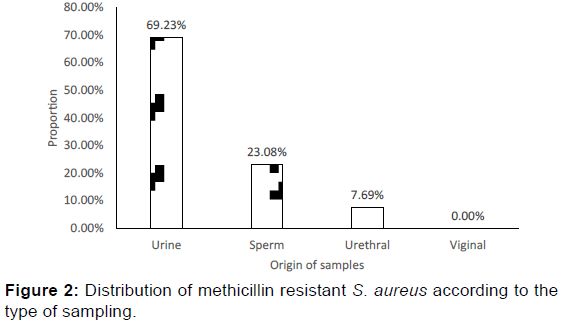
More than half (54.17%) of S. aureus strains were resistant to methicillin. Among the methicillin resistant S. aureus (MRSA), 69.23% were isolated from urine, 23.08% isolated from urethral and 07.69% from sperm [Figure 2]. Regarding the origin of MRSA, 92.31% of them were community-acquired.
Toxins productions
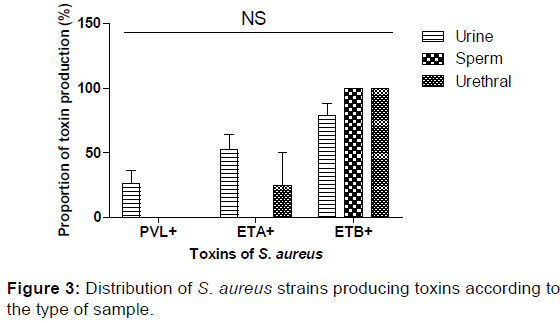
In our study, 26.32% of S. aureus strains isolated from the urine were LPV-producing; no strain of LPV-producing S. aureus was detected in urethral and sperm samples. Concerning the enterotoxins, ETB was produced by 83.33% of the S. aureus strains whereas about 45.33% produced ETA. Both ETA and ETB were produced by 33.33% of S. aureus strains. Regarding the type of sampling, ETA was produced by S. aureus isolated from urethral samples (25%) and urinary (52.63%). All the S. aureus strains isolated from the urethral and sperm samples produced ETB [Table 4]. The production of toxins by S. aureus was not a function of the type of sampling.
| Variables | MSSA (n=11) | MRSA (n=13) | P (Chi 2) | Significance | |
|---|---|---|---|---|---|
| PVL+ | 45.45% | 0% | 0.006 | *** | |
| ETA+ | 54.55% | 38.46% | 0.431 | NS | |
| ETB+ | 72.73% | 38.46% | 0.200 | NS | |
Table 4: Frequency of MRSA and MSSA producing LPV, ETA and ETB.
None of the MRSA strain was LPV-producing whereas 45.45% of MSSAs were LPV-producing. ETA was produced at different level by MSSA (54.55%) strains and of MRSA (38.46%), while ETB was highly produced MSAS (72.73%). The level of S. aureus producing PVL was a function of resistance to or not to methicillin [Figure 3].
Discussion
Among the analyzed vaginal samples, there was no pure or predominant culture of staphylococci. Previous studies reported that the main germs involve in the female genital infections are most often Chlamydiae trachomatis, Neisseria gonorrhoeae, Mycoplasma genitalium, Gardenella vaginalis and/or yeasts. [25] Staphylococcal strains of urinary origin were predominant (67.50%) and were derived from 61% of female patients, compared with 39% of male patients. Indeed, urine hasn’t any antimicrobial property, and can therefore be an excellent culture medium for bacteria growth. In addition, urinary tract infections (such as cystitis) are five times more common in women than in men because of the clothing habits and also of the proximity of the anal and vulvar regions. [14] The major part of the isolated staphylococci was coagulase-negative. This confirms the increasing role of coagulase-negative staphylococci (considered commensal) in staphylococcal infections. [16] In urinary infections, the predominant species in our study was S. saprophyticus (50%), which is comparable to the results obtained in Morocco. [13] In sperm, the coagulase-negative staphylococci were predominantly present with S. saprophyticus as the predominant species (61.91%). Thirty percent of the isolated strains were S. aureus mostly (35.19%) isolated from the urine. This proportion of S. aureus from urogenital origin is not far from the 38% reported in Morocco and Mauritania. [26] Probably due to the samples size and the adaptation of bacteria, the urine contamination rate we observed seems very higher that the 15.2% obtained in Benin by Baba-Moussa et al. [15]
The resistance profile of the staphylococci isolated strains varied according to the 19 antibiotics tested in our study [Table 3]. So, considering beta-lactams, penicillin G has been inactive on all Staphylococcus species. Methicillin resistance level varies from staphylococcal strains to another. Thus, all the S. haemolyticus and S. xylosus were resistant to that molecule whereas 54.17%, 52.50% and 38.46% of resistance were recorded respectively for S. aureus, S. saprophyticus, and S. epidermidis. Thus, to limit the presence of falsely positive methicillin resistance strains, it is useful to combine oxacillin to cefoxitin or latamoxef. [27] Amoxicillin and amoxicillin + clavulanic acid were also inactive on the five Staphylococcus species but with lower resistance levels for amoxicillin + clavulanic acid, and coagulase-negative staphylococci are naturally susceptible to beta-lactams. [28] In addition to beta-lactam antibiotics, four other antibiotics used fairly in Benin (ciprofloxacin, tetracycline, cotrimazole and doxycycline) were inactive on all five Staphylococcus species. However, staphylococci are reported to be naturally sensitive to fluoroquinolones. [29-31] A multi-resistance of S. aureus and coagulase-negative staphylococci observed in our study is consistent with that reported in other studies in Africa. [28,32] This result could be justified by the fact that in Cotonou, as elsewhere in Africa, several antibiotics molecules are available at affordable prices and are usually delivered without any medical prescription in pharmacies or are found in parallel markets for fake medicines. This sad reality promotes self-medication, over-consumption of antibiotics and consequently increased selection of resistance bacteria.
All the aminoglycosides used in our study were globally inactive on the five staphylococcal species and 83.33% of resistant strains to methicillin were of Kanamycin-Tobramycin-Gentamycin phenotype thought the coagulase-negative staphylococci strains are reported to be naturally sensitive to aminoglycosides. [29] This high resistance can be explained by the fact that aminoglycosides and gentamicin in particular constitute the antibiotic of choice for the treatment of severe staphylococcal infections and that are increasingly available at very affordable prices. Macrolides-Lincosamides-Streptogramines (MLS) were more active on four species (S. saprophyticus, S. epidermidis, S. xylosus and S. aureus). The constitutive MLSB phenotype was found in 28.75% of all staphylococci resistant to methicillin. Indeed, MLS, because of their high cost are still less accessible for the majority of population and this may explain the good activity of these antibiotics on our strains. The most active molecule on the Staphylococcus species identified during our study was nitrofurantoin with a resistance rate of 00% for S. saprophyticus, S. epidermidis, S. haemolyticus and 4.17% for S. aureus. Our data thus, confirms the efficacy of this antibiotic in the treatment of urinary and/or genital infections as previously reported. [33] Thus, patient are strongly invited not to use any molecules for the control of infection if not, the Staphylococcus involve in the urogenital infection will be more resistant. The reduction of the increase resistance among Staphylococcus in particular and globally all the bacterial strains must include all the actors (patient, practitioners, pharmacists and political authorities). The phenomenon is not only a burden for the patient and his family but also for the nation. Then, the populations must be made aware of the dangers resulting from their practices towards to the uses of drugs.
In our study, the rate of 54.17% of methicillin resistant S. aureus (MRSA) is close to the 50-57% reported in Senegal, [34] but significantly higher than the 1% observed in the Netherlands and Scandinavian countries, [35] and the 7.6% in Morocco. [26] Thus we can say that the frequency of MRSA varies greatly across countries. This remarkable resistance observed of S. aureus to methicillin can be explained by the consumption increasingly important and especially without medical opinion of this antibiotic by the population. This situation is not observed in the developed countries where the acquisition of any antimicrobial drug is certified by a medical prescription. Most (92.31%) of MRSA were isolated from the community. This rate is higher than the overall estimate of 30-40% found by some authors. [36] This is not surprising, as over the past two decades, cases of MRSA infections contracted outside the hospital have been regularly reported. [37] The high rate of community-acquired MRSA in Benin may be due to the dissemination of hospitalbased MRSA outside hospitals through inpatient (or companion) patients through the exchange of these strains within the community or during the home care. There a need to reduce this kind of contamination to reduce dissemination of resistance and pathogenic bacteria in the community. Methicillin resistance of the 13 strains of S. aureus was a crossed resistance with all betalactams, confirming previous work. [38,39] Approximately ninetytwo percent (92.31%) of these MRSA displays KTG phenotype. This is normal because more than 90% of MRSA are reported to be resistant to kanamycin, gentamicin and tobramycin. [28] In addition, about 40% of our MRSA were constitutive MLSB and 15.38% MLSB inducible. Nitrofurantoin was the only 100% active molecule on both SASM and MRSA and therefore remains an alternative in the treatment of urogenital infections.
Regarding the production of toxins, during our study, 26.32% of S. aureus strains all of urinary origin were PVL-producing. This rate is statistically different and slightly higher than the 21.4% found in Benin and France. [15] These S. aureus producers of PVL in our study were all community-based MSSA. This is not surprising because MSSAs (clonal complex CC121) LPV + are known to be common in many countries over the world. [40-42] Indeed, luk-PV genes can be incorporated into S. aureus lines by horizontal transfer, either before or after the acquisition of the mecA gene. [43,44] In this study, obtaining MSSA LPV + (45.45%) should draw attention to the existence of potential reservoirs of resistance and virulence factors that could lead to the emergence and spread of other MRSA LPV + in Benin. Our study showed that exfoliatins ETB (83.33%) was the most produced by S. aureus strains and then ETA (45.33%). This high production of exfoliatins, especially of ETB in urogenital affections, seems exceptional since exfoliatins release S. aureus toxins that are associated with well-established specific diseases (generalized exfoliation syndrome, bullous impetigo, etc.). [45] The question that may arise is whether exfoliatins are directly associated with urogenital infections or not.
Conclusion
Urinary-genital and particularly urinary infections are a growing concern in public health. In our study, a high proportion of coagulase-negative staphylococci were found. A high proportion of strains of multidrug-resistant staphylococci including betalactams, aminoglycosides, tetracycline, and cotrimazole were also obtained. This merely reflects the state of progress of the acquired resistance of these germs to antibiotics. Methicillinresistant Staphylococcus aureus was resistant to almost all antibiotics except lincomycin and pristinamycin (for some) and nitrofurantoin (for the whole), confirming the emergence in the community of strains of multi-resistant MRSA. No cases of LPV-producing MRSA (LPV +) strains were detected, but all were community-acquired strains of Staphylococcus aureus sensitive to methicillin and LPV-producing, potential reservoirs of MRSA LPV +.
Conflict of Interest
All authors disclose that there was no conflict of interest.
REFERENCES
- McCullough AR, Pollack AJ, Hansen MP, Glasziou PP, Looke DFM, Britt HC, et al. Antibiotics for acute respiratory infections in general practice: comparison of prescribing rates with guideline recommendations. Med. J. Aust. 2017; 207: 65-69.
- Cogen AL, Nizet V, Gallo RL. Skin microbiota: A source of disease or defense? The Br J Dermatol. 2008; 158: 442-455.
- Houssni B, Berkhli H, Madani H, Azzouzi A. Résistances bactériennes, consommation d'antibiotiques et politique de gestion de l'antibiothérapie. Officinal. 2011; 88: 1-3.
- Chancey ST, Zähner D, Stephens DS. Acquired inducible antimicrobial resistance in gram-positive bacteria. Future Microbiology. 2012; 7: 959-978.
- Munita JM, Arias CA. Mechanisms of antibiotic resistance. Microbiol Spectr. 2016; 4: 10.1128/microbiolspec.VMBF-0016-2015.
- Bæk KT, Gründling A, Mogensen RG, Thøgersen L, Petersen A, Paulander W, et al. ß-lactam Resistance in Methicillin-Resistant Staphylococcus aureus USA300 Is increased by inactivation of the ClpXP protease. Antimicrob Agents Chemother. 2014; 58: 4593-4603.
- Philippon A, Arlet G. Entérobactéries et bêta-lactamines : phénotypes de résistance naturelle. Pathologie Biologie. 2012; 60: 112-126.
- Rakotovao-Ravahatra ZD, Randriatsarafara FM, Rasoanandrasana S, Raverohanta L, Rakotovao AL. Phénotypes de résistance des souches d’Escherichia coli responsables d’infection urinaire au laboratoire du Centre Hospitalo-Universitaire de Befelatanana Antananarivo. Pan Afr. Med. J. 2017; 26:166.
- Li X, Sun J, Wu D, Wang L, Yang Y, Wang C, et al. Panton–Valentine leukocidin gene sequence variation and phage in methicillin-resistant and methicillin-susceptible Staphylococcus aureus from children in mainland China. Microbiol. Immunol. 2012; 56: 155–162.
- DeLeo FR, Chambers HF. Reemergence of antibiotic-resistant Staphylococcus aureus in the genomics era. J. Clin. Invest. 2009; 119: 2464-2474.
- Otto M. Staphylococcus aureus toxins. Cur. Opin. Microbiol. 2014; 0: 32-37.
- Tong SYC, Davis JS, Eichenberger E, Holland TL, Fowler VG. Staphylococcus aureus infections: Epidemiology, Pathophysiology, Clinical Manifestations, and Management. Clin Microbiol Rev. 2015; 28: 603-661.
- Sekhsokh Y, Chadli M, El-Hamzaoui SA. Frequency and antibiotic susceptibility of bacteria identified in urine. Med. Mal. Infect. 2008; 38: 324-327.
- Al-Badr A, Al-Shaikh G. Recurrent urinary tract infections management in women: A review. Sultan Qaboos Univ. Med. J. 2013; 13: 359-367.
- Baba-Moussa L, Anani L, Scheftel JM, Couturier M, Haïkou N, Hounsou F, et al. Virulence factors produced by strains of Staphylococcus aureus isolated from urinary tract infections. J. Hosp. Infect. 2008; 68: 32-38.
- Becker K, Heilmann C, Peters G. Coagulase-negative Staphylococci. Clin Microbiol Rev. 2014; 27: 870-926.
- Denis F, Bingen E, Martin C, Ploy MC, Quentin R. Bactériologie médicale Techniques usuelles. 2nd Edition Elsevier Masson, Paris, France, 2011; 640 p.
- Vandepitte J, Engbaek K, Piot P, Heuck CC. Bactériologie clinique: Techniques de base pour le laboratoire. Editions de l'OMS.1994; 25-110.
- Akoachere JFTK, Bughe RN, Oben BO, Ndip LM, Ndip RN. Phenotypic characterization of human pathogenic bacteria in fish from the coastal waters of South West Cameroon: Public health implications. Rev. Environ. Health. 2009; 24: 147-155.
- Cheesbrough M. District Laboratory Practice in Tropical Countries: Part 2. Cambridge University Press, Cambridge, UK. 2006; 299-329.
- SFM (Société Française de Microbiologie). Recommandations du Comité de l’Antibiogramme de la Société Française de Microbiologie. 2012; 59 p.
- Spaan AN, Henry T, Van Rooijen WJM, Perret M, Badiou C, Aerts CP, et al. The Staphylococcal toxin Panton-Valentine leukocidin targets human C5a receptors, Cell Host Microbe. 2013; 13: 584-594.
- Poutrel B. Validity of quantitative protein A, determination of "Staphylococcus aureus" by radial immunodiffusion. Ann. Microbiol. 1977; 128: 297-302.
- Aman MJ, Adhikari PR. Staphylococcal bicomponent pore-forming toxins: Targets for prophylaxis and immunotherapy. Toxins. 2014; 6: 950-972.
- Cu-Uvin S. Infectious diseases of the female genital tract, (5th edn). Clin. Infect. Dis. 2010; 50: 1076 p.
- Salem MLO, Ghaber SM, Baba SEWO, Maouloud MMO. Sensibilité aux antibiotiques des souches de Staphylococcus aureus communautaires dans la région de Nouakchott (Mauritanie). Pan Afr. Med. J. 2016; 24: 276.
- Bonjean M, Hodille E, Dumitrescu O, Dupieux C, Mongo NC, Allam C, et al. Disk diffusion testing for detection of methicillin-resistant Staphylococci: Does moxalactam improve upon cefoxitin? J. Clin. Microbiol. 2016; 54: 2905-2909.
- Ebongue CO, Tsiazok MD, Mefo’o JPN, Ngaba GP, Beyiha G, Adiogo D. Evolution de la résistance aux antibiotiques des entérobactéries isolées à l’Hôpital Général de Douala de 2005 à 2012. Pan Afr. Med. J. 2015; 20: 227.
- Gade ND, Qazi MS. Fluoroquinolone therapy in Staphylococcus aureus infections: Where do we stand? J. Lab. Physicians. 2013; 5: 109-112.
- Bispo PJM, Alfonso CE, Flynn WH, Miller D. Emerging 8-methoxyfluoroquinolone resistance among methicillin-susceptible Staphylococcus epidermidis isolates recovered from patients with Endophthalmitis. J. Clin. Microbiol. 2013; 51: 2959-2963.
- Randrianirina F, Soares JL, Ratsima E, Carod JF, Combe P, Grosjean P, et al. In vitro activities of 18 antimicrobial agents against Staphylococcus aureus isolates from the Institute Pasteur of Madagascar. Ann. Clin. Microbiol. Antimicrob. 2007; 6: 5.
- Kesah C, Ben Redjeb S, Odugbemi TO, Boye CS, Dosso M, Ndinya Achola JO, et al. Prevalence of methicillin resistant Staphylococcus aureus in eight African hospitals and Malta. Clin. Microbiol. Infect. 2003; 9: 153-156.
- Timothy OO, Olusesan FJ, Adesola BO, Temitayo AA, David FO, Ige OO. Antibiotic resistance pattern of bacterial isolates from cases of urinary tract infections among hospitalized and out-patients at a tertiary health facility in South Western Nigeria. Ann Trop Med. Public Health. 2014; 7: 130-135.
- Seydi M, Sow AI, Soumaré M, Diallo HM, Hatim B, Tine RCK, et al. Place des bactériémies à Staphylococcus aureus au CHU de Fann à Daka. Med. Mal. Infect. 2004; 34: 210-215.
- Green BN, Johnson CD, Egan JT, Rosenthal M, Griffith EA, Evans MW. Methicillin-resistant Staphylococcus aureus: an overview for manual therapists. J. Chiropr. Med. 2012; 11: 64-76.
- Udo EE. Community-acquired methicillin-resistant Staphylococcus aureus: The new face of an old foe? Med. Princ. Pract. 2013; 22: 20-29.
- Sobhy N, Aly F, Abd El-Kader Ola, Ghazal A, Elbaradei A. Community-acquired methicillin-resistant Staphylococcus aureus from skin and soft tissue infections (in a sample of Egyptian population): analysis of mec gene and Staphylococcal cassette chromosome. Braz. J. Infect. Dis. 2012; 16: 426-431.
- Chambers HF. Methicillin resistance in staphylococci: Molecular and biochemical basis and clinical implications. Clin. Microbiol. Rev. 1997; 10: 781-791,.
- Brzychczy-Wloch M, Borszewska-Kornacka M, Gulczynska E, Wojkowska-Mach J, Sulik M, Grzebyk M, et al. Prevalence of antibiotic resistance in multi-drug resistant coagulase-negative staphylococci isolated from invasive infection in very low birth weight neonates in two Polish NICUs. Ann. Clin. Microbiol. Antimicrob. 2013; 12: 41.
- Rasigade JP, Laurent F, Lina G, Meugnier H, Bes M, Vandenesch F, et al. Global distribution and evolution of Panton-Valentine leukocidin-positive methicillin-susceptible Staphylococcus aureus, 1981-2007. J. Infect. Dis. 2010; 201: 1589-1597.
- Schaumburg F, Alabi AS, Peters G, Becker K. New epidemiology of Staphylococcus aureus infection in Africa. Clin. Microbiol. Infect. 2014; 20: 589-596.
- Shittu AO, Okon K, Adesida S, Oyedara O, Witte W, Strommenger B, et al. Antibiotic resistance and molecular epidemiology of Staphylococcus aureus in Nigeria. BMC Microbiol. 2011; 11: 92.
- Jung J, Song EH, Park SY, Lee SR, Park SJ, Sung H, et al. Emergence of Panton-Valentine leucocidin-positive ST8-methicillin-resistant Staphylococcus aureus (USA300 clone) in Korea causing healthcare-associated and hospital-acquired bacteraemia. Eur J Clin Microbiol Infect Dis. 2016; 35: 1323-1329.
- Alibayov B, Baba-Moussa L, Sina H, Zdenková K, Demnerová K. Staphylococcus aureus mobile genetic elements. Mol. Biol. Rep. 2014; 41: 5005-5018.
- Nordenfelt P, Collin M. Bacterial pathogenesis: Methods and protocols. Meth Mol Biol. 2017; 1535: 357p.



 The Annals of Medical and Health Sciences Research is a monthly multidisciplinary medical journal.
The Annals of Medical and Health Sciences Research is a monthly multidisciplinary medical journal.