Body Mass Index in Pregnancy Does Not Affect Peroxisome Proliferator‑activated Receptor Gamma Promoter Region (−359 to −260) Methylation in the Neonate
- *Corresponding Author:
- Dr. Hugo Mendieta Zerón
Faculty of Medicine, Autonomous University of the State of Mexico (UAEMex), Felipe Villanueva sur 1209, Col. Rancho Dolores CP, 50170 Toluca, Mexico.
E-mail: mezh_74@yahoo.com
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms.
Citation Casamadrid V, Amaya CA, Mendieta ZH. Body mass index in pregnancy does not affect peroxisome proliferator-activated receptor gamma promoter region (-359 to -260) methylation in the neonate. Ann Med Health Sci Res 2016;6:38-43.
Abstract
Background: Obesity in pregnancy can contribute to epigenetic changes. Aim: To assess whether body mass index (BMI) in pregnancy is associated with changes in the methylation of the peroxisome proliferator‑activated receptor γ (PPAR) promoter region (−359 to − 260) in maternal and neonatal leukocytes. Subjects and Methods: In this matched, cohort study 41 pregnant women were allocated into two groups: (a) Normal weight (n = 21) and (b) overweight (n = 20). DNA was extracted from maternal and neonatal leukocytes (4000–10,000 cells) in MagNA Pure (Roche) using MagNA Pure LC DNA Isolation Kit 1 (Roche, Germany). Treatment of DNA (2 μg) was performed with sodium bisulfite (EZ DNA Methylation‑Direct™ Kit; Zymo Research). Real‑time quantitative polymerase chain reaction (qPCR) was performed in a LightCycler 2.0 (Roche) using the SYBR® Advantage® qPCR Premix Kit (Clontech). The primers used for PPARγ coactivator (PPARG) M3 were 5’‑aagacggtttggtcgatc‑3’ (forward), and5’‑cgaaaaaaaatccgaaatttaa‑3’ (reverse) and those for PPARG unmethylated were: 5’‑gggaagatggtttggttgatt‑3’ (forward) and 5’‑ttccaaaaaaaaatccaaaatttaa‑3’ (reverse). Intergroup differences were calculated using the Mann–Whitney U‑test, and intragroup differences, with the Wilcoxon test (IBM SPSS Statistics for Windows, Version 19.0. Armonk, NY: IBM Corp.). Results: Significant differences were found in BMI, pregestational weight, and postdelivery weight between groups but not in the methylation status of the PPARγ promoter region (−359 to − 260). Conclusion: The PPARγ promoter region (−359 to − 260) in peripheral leukocytes is unlikely to get an obesity‑induced methylation in pregnancy.
Keywords
Body mass index, Methylation, Peroxisome proliferator-activated receptor gamma, Pregnancy
Introduction
Obesity in pregnancy has harmful effects on maternal health.[1,2] Moreover, the mother’s increased adiposity has been suggested to influence the programming of metabolic pathways in the fetus, predisposing it to cardiovascular disease and diabetes[3,4] due to epigenetic mechanisms.[5]
A newly recognized primary cause of obesity epidemic is the developmental programming effects of normal or excessive weight newborns exposed to maternal obesity and high-fat diets.[6] During embryonic development, an appropriate DNA methylation state should be maintained during the rapid step cycles of cell proliferation.[7] It has been postulated that maternal obesity may impair DNA methylation of imprinted genes,[8] but the detailed mechanisms underlying these changes remain unknown. Other studies have shown that the expression of peroxisome proliferator-activated receptor a (PPAR) is regulated by DNA methylation in its promoters.[9,10] Following this line, PPAR? is a member of the hormonal nuclear receptor superfamily[11] with four known isoforms.[12] PPAR?1 and-?2 are expressed predominantly in adipocytes,[13] and the former is additionally expressed in breast, colon, vascular cells and in small amounts, in the heart and liver.[14] PPAR?3 and-?4 have been identified in macrophages, and PPAR?3 has been found in colon.[12]
PPAR? participates in the differentiation of preadipocytes into adipocytes,[15] meaning that a permanent activation of this gene could explain a higher incidence of obesity at lower age in children born to obese women. Persistent alteration in mRNA expression of PPAR? promoter region (-359 to -260) methylation has been associated with an increased risk of colon cancer,[16] hyperhomocysteinemia,[17] liver inflammation, and fibrosis in patients with hepatitis B,[18] etc. Our main objective was to assess whether clinical, anthropometric, and biochemical variables of the mother were associated with changes in the methylation of this PPAR? promoter region in maternal and neonatal leukocytes.
Subjects and Methods
Ethical clearance was granted by the Research Committee of the Maternal-Perinatal Hospital “Mónica Pretelini Sáenz” (HMPMPS) (code: 05-06-2009) and followed the General Health Research Law of Mexico, and the Declaration of Helsinki (Fortaleza, Brazil). All patients were asked to sign written informed consent.
Study site and design
This was a matched cohort study conducted from July 2009 to September 2012, at the tertiary health level HMPMPS, Health Institute of the State of Mexico, Toluca, Mexico.
Study population and ethical issue
Pregnant women were recruited in the first trimester of pregnancy and did not include cases with congenital heart and disabling or autoimmune diseases, this information was obtained from medical records in the first obstetrical visit. Those whose clinical follow-ups were lost or who, if in the postpartum required attention at the Obstetric Intensive Care Unit were eliminated from the study (n = 2).
Sample size
Accepting an alpha risk of 0.05 and a beta risk of 0.2 in a two-sided test, twenty subjects per group were necessary to find as statistically significant a proportion difference in the methylation status of the PPAR? promoter region (-359 to -260), expected to be of 0% in pregnant women with normal weight (NW) and 40% in those with overweight (OW).
Data collection techniques and tools
One clinical visit per month was established. Body weight and height were measured during an overnight fasting state using an adult scale (Seca, Hamburg, Germany). Prepregnancy body mass index (BMI) was calculated as weight in kg divided by height in square meters based on the prenatal chart or on the self-reported weight of women with no prenatal chart.
Blood pressure was recorded at each visit using a standard sphygmomanometer (Riester Big Ben® Square; Jangingen, Germany). Preeclampsia was diagnosed and classified according to the American College of Obstetricians and Gynecologists guidelines.
Fasting blood samples (10 ml) were taken at the HMPMP Laboratory at an early-morning appointment after an overnight fast of 8 h. Serum samples were analyzed for glucose and lipid profile (Dimension RxL Max™; Dade Behring, USA). At the end of pregnancy, 1?2 ml of a neonatal peripheral blood sample for leukocyte extraction was taken.
Dietetic treatment was calculated according to the height, weeks of gestation, and weight, considering an energy intake of 30 kcal/kg of ideal weight and a macronutrient distribution of 55?65% carbohydrates, 10?20% fat, and the remainder as proteins. On each nutritional appointment, the Healthy Eating Index for Pregnancy (HEI) was evaluated and all women were recommended to include methionine-rich foods (beans, eggs, fish, garlic, lentils, onion, and soy) and those containing folic acid and Vitamin B12 (beef liver, cereals, whole grains, yeast, etc.) in adequate quantities in their diet. The information was complemented with the Food Frequency Questionnaire, and adherence to the diet was considered adequate with 80% compliance to the indicated calories, on at least four appointments.
Leukocytes were obtained according to the ACK lysing buffer (Lonza) protocol.[19] Briefly, a peripheral blood sample was placed in an EDTA tube and then centrifuged at 2500 rpm for 10 min. All samples were maintained at -80°C until further analysis.
DNA was extracted from maternal and neonatal leukocytes (4000?10,000 cells) in the Magna Pure LC 2.0 Instrument (Roche, Germany) using MagNA Pure LC DNA Isolation Kit 1 (Roche, Germany).[20] After extraction, the DNA was quantified using a NanoPhotometer (Implen GmbH,Germany), reporting concentration (in µg/ml) and purity (as 260/280 absorbance).
Treatment of DNA (2 µg) was performed with sodium bisulfite (EZ DNA Methylation-Direct™ Kit; Zymo Research). For the control group, we used purified human methylated and unmethylated (U3) DNA (Zymo Research) with specific oligonucleotides. Lymphocyte DNA from healthy nonpregnant donors was used as negative control and DNA methylated in vitro with the Sss I enzyme (New England BioLabs) was used as positive control for methylation.
The methylated (M3) and U3 primers employed were those proposed by Pancione et al.[16] These primers comprise the same region, that is, -359 to - 260, with respect to the transcription site, but differ in sequence with respect to cytosines of the CpG islands. The M3 primer set is specific to the case in which CpG cytosines are methylated, thus not modified, during the bisulfite treatment, whereas the U3 primer set is specific for the opposite case, that is, in which all cytosines are modified during bisulfite treatment.
Primer functionality was verified utilizing the following methods: First, by analyzing the primer sequences based on the in silico change of the gene sequence obtained from PubMed (NC_000003.11), and second, by obtaining theoretical products and running primer tests of specificity in the methBLAST sequence similarity search program.
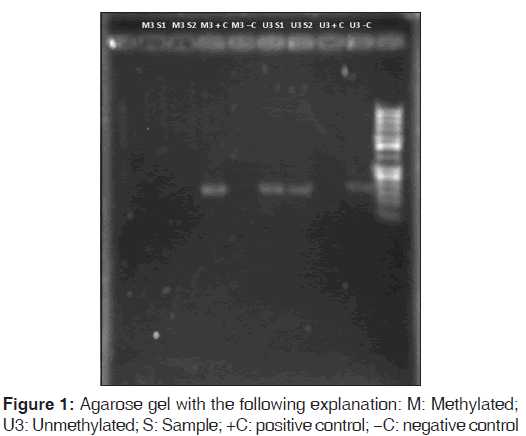
In the polymerase chain reaction (PCR) analysis the primers employed for PPARg coactivator (PPARG) M3 were: 5’-aagacggtttggtcgatc-3’ (forward) and 5’-cgaaaaaaaatccgaaatttaa-3’ (reverse) and those for PPARG U3 were: 5’-gggaagatggtttggttgatt-3’ (forward) and 5’-ttccaaaaaaaaatccaaaatttaa-3’ (reverse). In a life express thermal cycler (Bioer, China), the steps were as follows: (1) denaturation: 90°C for 15 s, (2) amplification (32 cycles): 95°C 5 s, 53°C 20 s, 72°C 30 s, and 76°C 30 s. DNA integrity and methylation status were visualized in 2% agarose gels, stained with ethidium bromide, and visualized under an ultraviolet transilluminator (Gel Logic 212 Pro, Carestream, USA). Finally, PCR products were sequenced and read with Chromas software version 2.33 (Instituto de Neurobiología, National Autonomous University of Mexico, UNAM, Juriquilla, Querétaro, Mexico) to verify correct primer amplification.
Data analysis
Statistical analysis was performed using SPSS version 19 (IBM SPSS Statistics for Windows, Version 19.0. Armonk, NY: IBM Corp.). First, descriptive analyses were performed; then, we estimated the differences between the study groups using the Mann–Whitney U-test or the Student’s t-test depending on the Kolmogorov test to determine the distribution of the data. Finally, through the paired sample t-test or the Wilcoxon test we compared whether the same group presented differences across time. Difference was considered significant at P = 0.05.
Results
Forty-one pregnant women were allocated into two groups: (a) NW pregnant women (n = 21) and their offsprings and (b) OW pregnant women (n = 20) and their offsprings. The anthropometric variables of the pregnant women are listed in Table 1. The weight of the newborns was 2999 (381) g and 2958 (616) g for those of NW and OW women, respectively, and infant length was 49.55 (2.13) cm and 49.32 (3.06) cm for those of NW and OW women, respectively.
| Variable | Group | P | |
|---|---|---|---|
| Normal weightmean (SD) | Overweightmean (SD) | ||
| Age (years) | 23.9 (5.4) | 25.9 (6.9) | |
| PGW (kg) | 54.5 (6.7) | 68.1 (10) | <0.001 |
| Height (cm) | 158.5 (6.8) | 154.7 (3.9) | |
| BMI (kg/m2)* | 21.6 (1.9) | 28.5 (3.8) | <0.001 |
| PDW (kg) | 62.4 (6.4) | 70.7 (9.9) | <0.001 |
| BMI (kg/m2)† | 24.8 (2.1) | 29.5 (3) | <0.001 |
| Weight gain (kg) | 11.1 (3.2) | 9.0 (6.2) | |
Table 1: Anthropometric variables by group
On analyzing all of the data, we found no statistical differences in age, pregnancy duration, or net weight gain, but significant differences were found in BMI, pregestational weight, and postdelivery weight between groups.
In blood pressure, no statistical differences could be established with the Friedman test, except for the third-trimester pregnancy for the OW group. We also performed a similar analysis among trimesters, comparing both groups and obtaining the same unique exception (data not shown). Only one confirmed case of preeclampsia occurred within the OW group. This patient exhibited outstanding cholesterol and triglyceride values.
Table 2 summarizes intragroup statistical results compared by trimester, demonstrating the following cases: (a) Variables with which evolution presented no statistical differences (this is the case of glucose in the NW group and of high-density lipoprotein cholesterol in both groups), (b) one variable with an increase from the first to the second trimester, followed by stagnation (glucose in the OW group), (c) variables that increased continuously (cholesterol in the NW group and triglycerides in both groups), (d) variables that showed differences in relation to the first trimester (cholesterol in the OW group, low-density lipoproteins cholesterol in both groups, and weight-gain in the OW group), and (e) one variable with an increase from the second to the third trimester (weight gain in the OW group). From these data, we can observe that the majority of the variables had a significant statistical difference when compared against the first trimester (eight cases with the second trimester and eight cases with the third trimester). As for the intergroup analysis (NW vs. OW), we found no statistical differences, except for weight gain during the first-trimester visit, 2.3 (2.2) kg versus - 0.38 (3) kg, respectively (P < 0.01).
| Variable | Group | |||||
|---|---|---|---|---|---|---|
| NW trimester | OW trimester | |||||
| 1 | 2 | 3 | 1 | 2 | 3 | |
| Cholesterol (mg/dL) | 173 (36) | 216 (46) | 249 (63) | 175(21) | 219 (47) | 232 (49) |
| P | <0.001x | <0.001y | <0.001z | <0.001x | 0.09y | <0.001z |
| Glucose (mg/dL) | 84 (12) | 78 (8) | 76 (7) | 82 (5) | 77 (6) | 78 (7) |
| P | 0.07x | 0.25y | 0.06z | <0.01x | 0.51y | 0.05z |
| HDL-C (mg/dL)† | 62 (28) | 64 (16) | 62 (14) | 52 (8) | 60 (16) | 54 (12) |
| P | 0.07x | 0.44y | 0.144z | 0.05x | 0.11y | 0.37z |
| LDL-C (mg/dL) | 101 (33) | 130 (38) | 145 (54) | 102 (20) | 131 (39) | 146 (43) |
| P | <0.001x | <0.09y | <0.001z | <0.001x | 0.07y | <0.001z |
| Triglycerides (mg/dL) | 136 (56) | 201 (75) | 247 (60) | 147 (51) | 215 (67) | 255 (65) |
| P | <0.001x | <0.01y | <0.001z | <0.001x | <0.01y | <0.001z |
| Weight gain (kg) | 2.3 (2.2) | 3.8 (3.1) | 4.7 (2.6) | -0.38 (3) | 3.6 (3.5) | 5.7 (3.9) |
| P | 0.11x | 0.38y | <0.01z | <0.01x | 0.09y | <0.001z |
Table 2: Biochemical variables grouped by trimester*
Taking into account the pregnancy weight-gain recommendations of 11.5?16 kg for NW and of 7?11.5 kg for the OW case,[21] the percentage of pregnant women who maintained an ideal weight gain was 33.3% (7/21) in the NW and 40% (8/22) in the OW group. According to HEI, there was adequate adherence to nutritional counseling in 38.1% (8/21) of women in the NW group and in 25% (5/20) of women in the OW group.
All maternal and neonatal peripheral leukocyte samples were found to be U3 in both groups. These results were verified by both quantitative PCR amplification with the unmethylation-specific primer and the absence of amplification in the presence of the methylation-specific primer [Figure 1].
Discussion
The methylation status of g (PPARGC1A and PPARg genes has shown to be important for the study of metabolic programming, and a role has been postulated of the BMI in this effect. In our study with pregnant women, the compared groups differed in weight-related variables, with essentially no statistical differences for all remaining variables. This is an appreciable characteristic because it limits any other difference aside from the effects of BMI. Other than BMI, both groups had differences in postpartum weight. When recalculating BMI after delivery, the initial NW group was found within the classification of OW, and the initial OW group fell into the obesity grade I range. The NW group was noted as having a higher net weight gain because recommendations are more extensive in these women.
Surprisingly, when comparing biochemical variables between groups among the trimesters, no statistical difference was found among these although it must be clarified that the cohort groups followed different trends and the intragroup difference for cholesterol and triglycerides from the first to the second and third trimester in each group could be associated to the BMI increase. In this behavior, pregnant women who adhered to the nutritional guidance afforded exhibited a slight decrease in weight gain, probably due to the influence of the dietetic orientation. In analyzing adherence to nutritional guidance, this action apparently contributed to the moderate weight gain, probably because of the influence of the diet orientation.
Regarding gestation weeks and the weight and length of the newborn at birth, no differences were found. While Grados Valderrama et al.[22] found a positive correlation between newborn weight and maternal BMI and weight gain, we found no correlation, probably because of the reduced study group size.
In pregnancy, it is normal to have hyperlipidemia.[23,24] Indeed, the increase in cholesterol is necessary because it is the precursor of Vitamin D[25] and corticosteroid hormones, both critical for a healthy pregnancy.[26] In this study, we observed an increase of cholesterol in both groups that was higher in the third quarter of pregnancy. High cholesterol levels in pregnancy were believed to actually give rise to fetal programming for atherogenesis.[27]
Solomon et al.[28] found that pregnant women with high cholesterol levels and elevated insulin levels were more likely to develop pregnancy-induced hypertension or preeclampsia.
In our study, although cholesterol and triglycerides increased in both groups, the increase was more noticeable in triglycerides. As published previously, triglyceride levels increase up to three times as long as the pregnancy progresses through the third quarter.[5] Similar results were obtained in this study, suggesting changes in lipid metabolism, which may be accompanied by functional and morphological changes in adipocytes.
In our approach, we found no different methylation index in any case. In fact, all samples were found to be U3. Previous efforts have been performed to analyze the methylation changes in several genes, including PPAR?, with contrasting results.[29,30] Data from Jacoby et al.[31] suggest that inter-individual variability and co-regulation of DNA methylation differ among blood cell populations. It is noteworthy that each author might choose a different region of the same gene to be analyzed. The diet program during pregnancy should also be considered because strict diet supervision guaranteed adequate portions of methionine, folic acid, and Vitamin B12, all with a known role in preventing methylation disequilibrium.
The available information is overwhelming, in showing that the PPAR?2 promoter is particularly sensitive to nutritional changes.[32,33] Even more, animal models have shown that offspring of diet-induced obese dams have altered mRNA expression of PPAR?.[34,35] Thus, methylation status is one of the epigenetic changes that can explain maternal programming of metabolic syndrome-related phenotypes,[36] but further work is required to define the role of PPAR?.
It cannot be ruled out that the enzymatic activity in each tissue is of paramount importance because the profile of methylation observed in leukocytes might be different from fat tissue, which is directly involved in metabolic balance and obesity.
Finally, despite the great efforts invested in explaining some possible factors that predispose to obesity genetically and epigenetically, a single locus has not yet been identified as being entirely responsible for this pathology. Whether epigenetic reprogramming that operates during oocyte formation and in the initial stages of embryogenesis[37] is involved in the obesity transgenerational inheritance has not been clarified. The current trend, given the advances in molecular biology presented in the last few years, is to search for Genome-wide associations, which can provide a better understanding of the pathology of obesity and transgenerational health consequences.[38]
A limitation of this study is the evaluation of a sole promoter region, but the negative results in mothers and babies for a methylated status points out the relevance of evaluating other PPAR? promoter regions that could be useful in the maternal programming of metabolic diseases on the fetus. A significant edge of our work is the fact that while most of the studies related to the PPAR methylation have been performed in solid organ tissues,[39,40] the analysis in neonatal blood, as we did, is scarce.[30]
Conclusion
While further work is required to define in detail the epigenetic changes induced by obesity in pregnancy, our results show that the PPAR? promoter region (-359 to - 260) is unlikely to be easily methylated in peripheral leukocytes in this physiological weight gain condition.
Acknowledgements
Authors thank the National Council of Science and Technology (CONACYT), México, for the MSc. Scholarship awarded to Ruth Elizabeth Casamadrid Vázquez and Maggie Brunner M.A., for her excellent help with the English style correction.
Financial support and sponsorship
National Council of Science and Technology (CONACYT), Mexico and Asociación Científica Latina A.C (ASCILA).
Conflicts of interest
There are no conflicts of interest.
References
- English FA, Kenny LC, McCarthy FP. Risk factors and effective management of preeclampsia. Integr Blood Press Control 2015;8:7-12.
- Lim CC, Mahmood T. Obesity in pregnancy. Best Pract Res Clin Obstet Gynaecol 2015;29:309-19.
- Pereira TJ, Moyce BL, Kereliuk SM, Dolinsky VW. Influence of maternal overnutrition and gestational diabetes on the programming of metabolic health outcomes in the offspring: Experimental evidence. Biochem Cell Biol 2014:1-14.
- Li CC, Maloney CA, Cropley JE, Suter CM. Epigenetic programming by maternal nutrition: Shaping future generations. Epigenomics 2010;2:539-49.
- Barua S, Junaid MA. Lifestyle, pregnancy and epigenetic effects. Epigenomics 2015;7:85-102.
- Desai M, Ross MG. Fetal programming of adipose tissue: Effects of intrauterine growth restriction and maternal obesity/high-fat diet. Semin Reprod Med 2011;29:237-45.
- Reik W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 2007;447:425-32.
- Howie GJ, Sloboda DM, Kamal T, Vickers MH. Maternal nutritional history predicts obesity in adult offspring independent of postnatal diet. J Physiol 2009;587(Pt 4):905-15.
- Burdge GC, Lillycrop KA, Phillips ES, Slater-Jefferies JL, Jackson AA, Hanson MA. Folic acid supplementation during the juvenile-pubertal period in rats modifies the phenotype and epigenotype induced by prenatal nutrition. J Nutr 2009;139:1054-60.
- Slater-Jefferies JL, Lillycrop KA, Townsend PA, Torrens C, Hoile SP, Hanson MA, et al. Feeding a protein-restricted diet during pregnancy induces altered epigenetic regulation of peroxisomal proliferator-activated receptor-a in the heart of the offspring. J Dev Orig Health Dis 2011;2:250-5.
- Lazar MA. PPAR?, 10 years later. Biochimie 2005;87:9-13.
- Arck P, Toth B, Pestka A, Jeschke U. Nuclear receptors of the peroxisome proliferator-activated receptor (PPAR) family in gestational diabetes: From animal models to clinical trials. Biol Reprod 2010;83:168-76.
- Janani C, Ranjitha Kumari BD. PPAR gamma gene – A review. Diabetes Metab Syndr 2015;9:46-50.
- Vidal-Puig AJ, Considine RV, Jimenez-Liñan M, Werman A, Pories WJ, Caro JF, et al. Peroxisome proliferator-activated receptor gene expression in human tissues. Effects of obesity, weight loss, and regulation by insulin and glucocorticoids. J Clin Invest 1997;99:2416-22.
- Lecoutre S, Breton C. Maternal nutritional manipulations program adipose tissue dysfunction in offspring. Front Physiol 2015;6:158.
- Pancione M, Sabatino L, Fucci A, Carafa V, Nebbioso A, Forte N, et al. Epigenetic silencing of peroxisome proliferator-activatedreceptor ? is a biomarker for colorectal cancer progression and adverse patients’ outcome. PLoS One 2010;5:e14229.
- Yideng J, Zhihong L, Jiantuan X, Jun C, Guizhong L, Shuren W. Homocysteine-mediated PPARalpha, gamma DNA methylation and its potential pathogenic mechanism in monocytes. DNA Cell Biol 2008;27:143-50.
- Zhao Q, Fan YC, Zhao J, Gao S, Zhao ZH, Wang K. DNA methylation patterns of peroxisome proliferator-activated receptor gamma gene associated with liver fibrosis and inflammation in chronic hepatitis B. J Viral Hepat 2013;20:430-7.
- Available from: http://www.bio.lonza.com/uploads/ tx_mwaxmarketingmaterial/Lonza_Manuals Product Instructions_ACK_Lysing_Buffer.pdf. [Last accessed on 2016 Jan 30].
- Available from: https://www.lifescience.roche.com/shop/ products/magna-pure-lc-dna-isolation-kit-i. [Last accessed on 2016 Jan 30].
- Rasmussen KM, Yaktine AL, editors. Weight Gain During Pregnancy: Reexamining the Guidelines. Washington. D.C., USA: National Academies Press; 2009. p. 324.
- Grados Valderrama FM ,Cabrera Epiquén R , Díaz Herrera J. [Prepregnancy nutritional status and maternal weight gain during pregnancy and its relation to birth weight]. Rev Med Hered 2003;14:128-33.
- Garduño-Alanís A, Vázquez-de Anda G, Valdés-Ramos R, Talavera JO, Herrera-Villalobos JE, Huitrón-Bravo GG, et al. Predictors of hyperlipidemia during the first half of pregnancy in Mexican women. Nutr Hosp 2014;31:508-13.
- Mendieta-Zerón H, Huerta-Coyote O. Dyslipidemia is a persistent problem in puerperium with or without preeclampsia. Clin Exp Obstet Gynecol 2013;40:229-32.
- Bartels A, O’Donoghue K. Cholesterol in pregnancy: A review of knowns and unknowns. Obstet Med 2011;4:147-51.
- Kuzawa CW, Adair LS. Lipid profiles in adolescent Filipinos: Relation to birth weight and maternal energy status during pregnancy. Am J Clin Nutr 2003;77:960-6.
- Szostak-Wegierek D. Intrauterine nutrition: Long-term consequences for vascular health. Int J Womens Health 2014;6:647-56.
- Solomon CG, Carroll JS, Okamura K, Graves SW, Seely EW. Higher cholesterol and insulin levels in pregnancy are associated with increased risk for pregnancy-induced hypertension. Am J Hypertens 1999;12:276-82.
- Gemma C, Sookoian S, Alvariñas J, García SI, Quintana L, Kanevsky D, et al. Maternal pregestational BMI is associated with methylation of the PPARGC1A promoter in newborns. Obesity (Silver Spring) 2009;17:1032-9.
- Sharp GC, Lawlor DA, Richmond RC, Fraser A, Simpkin A, Suderman M, et al. Maternal pre-pregnancy BMI and gestational weight gain, offspring DNA methylation and later offspring adiposity: Findings from the Avon longitudinal study of parents and children. Int J Epidemiol 2015;44:1288-304.
- Jacoby M, Gohrbandt S, Clausse V, Brons NH, Muller CP. Interindividual variability and co-regulation of DNA methylation differ among blood cell populations. Epigenetics 2012;7:1421-34.
- Yamazaki T, Shiraishi S, Kishimoto K, Miura S, Ezaki O. An increase in liver PPARgamma2 is an initial event to induce fatty liver in response to a diet high in butter: PPARgamma2 knockdown improves fatty liver induced by high-saturated fat. J Nutr Biochem 2011;22:543-53.
- Schadinger SE, Bucher NL, Schreiber BM, Farmer SR. PPARgamma2 regulates lipogenesis and lipid accumulation in steatotic hepatocytes. Am J Physiol Endocrinol Metab 2005;288:E1195-205.
- Samuelsson AM, Matthews PA, Argenton M, Christie MR, McConnell JM, Jansen EH, et al. Diet-induced obesity in female mice leads to offspring hyperphagia, adiposity, hypertension, and insulin resistance: A novel murine model of developmental programming. Hypertension 2008;51:383-92.
- Bayol SA, Simbi BH, Bertrand JA, Stickland NC. Offspring from mothers fed a ‘junk food’ diet in pregnancy and lactation exhibit exacerbated adiposity that is more pronounced in females. J Physiol 2008;586:3219-30.
- Burgueño AL, Cabrerizo R, Gonzales Mansilla N, Sookoian S, Pirola CJ. Maternal high-fat intake during pregnancy programs metabolic-syndrome-related phenotypes through liver mitochondrial DNA copy number and transcriptional activity of liver PPARGC1A. J Nutr Biochem 2013;24:6-13.
- Iovino N. Drosophila epigenome reorganization during oocyte differentiation and early embryogenesis. Brief Funct Genomics 2014;13:246-53.
- Herrera BM, Keildson S, Lindgren CM. Genetics and epigenetics of obesity. Maturitas 2011;69:41-9.
- Laker RC, Lillard TS, Okutsu M, Zhang M, Hoehn KL, Connelly JJ, et al. Exercise prevents maternal high-fat diet-induced hypermethylation of the Pgc-1a gene and age-dependent metabolic dysfunction in the offspring. Diabetes 2014;63:1605-11.
- Barrès R, Yan J, Egan B, Treebak JT, Rasmussen M, Fritz T, et al. Acute exercise remodels promoter methylation in humanskeletal muscle. Cell Metab 2012;15:405-11.




 The Annals of Medical and Health Sciences Research is a monthly multidisciplinary medical journal.
The Annals of Medical and Health Sciences Research is a monthly multidisciplinary medical journal.