Emergence of Multidrug Resistance and Metallo‑beta‑lactamase Producing Acinetobacter baumannii Isolated from Patients in Shiraz, Iran
- *Corresponding Author:
- Prof. Motamedifar M,
Department of Bacteriology and Virology, School of Medicine, Shiraz University of Medical Sciences, Zand St, Imam Hossein Sq, Shiraz, Iran.
E-mail: motamedm@sums.ac.ir
Citation: Moghadam MN, Motamedifar M, Sarvari J, Sedigh ESH, Mousavi SM, Moghadam FN. Emergence of multidrug resistance and metallo-beta-lactamase producing Acinetobacter baumannii isolated from patients in Shiraz, Iran. Ann Med Health Sci Res 2016;6:162-7.
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms.
Abstract
Background: MetalloĂƒÂ¢Ă‚â‚¬Ă‚â€˜betaĂƒÂ¢Ă‚â‚¬Ă‚â€˜lactamase (MβL) enzymes production is one of the most important resistance mechanisms against carbapenems in some bacteria including Acinetobacter baumannii. Aims: This study was aimed to determine the antimicrobial susceptibility and the prevalence of MβL among carbapenemĂƒÂ¢Ă‚â‚¬Ă‚â€˜resistant isolates of A. baumannii. Materials and Methods: In this crossĂƒÂ¢Ă‚â‚¬Ă‚â€˜sectional study from October 2012 to April 2013, 98 isolates were identified as A. baumannii using Microgen™ kits and confirmed by molecular method. These isolates were tested for antimicrobial susceptibilities by disk diffusion method according to the Clinical and Laboratory Standards Institute guidelines. CarbapenemĂƒÂ¢Ă‚â‚¬Ă‚â€˜resistant isolates were further detected phenotypically by MβL minimal inhibitory concentration (MIC)ĂƒÂ¢Ă‚â‚¬Ă‚â€˜test strips, and subsequently positive MβL isolates were confirmed by polymerase chain reaction (PCR). Results: Overall, 98% (96/98) of A. baumannii isolates were detected as carbapenemĂƒÂ¢Ă‚â‚¬Ă‚â€˜resistant by MIC test. Highest sensitivity to the tested antibiotic with 42.9% (42/98) was observed to colistin. Of 96 carbapenemĂƒÂ¢Ă‚â‚¬Ă‚â€˜resistant isolates, 43 were phenotypically positive for MβL; out of 43 isolates, 37 were confirmed for the presence of MβL genes by PCR. Conclusion: The frequency of drug resistance among the clinical samples of A. baumannii isolated in our study against most of the antibiotics was very high. Moreover, all MβL producing isolates were multidrug resistance. Therefore, systematic surveillance to detect MβL producing bacteria and rational prescription and use of carbapenems could be helpful to prevent the spread of carbapenem resistance.
Keywords
Acinetobacter baumannii, Antibiotic resistance, Carbapenem, Iran, Metallo-beta-lactamase
Introduction
Acinetobacter baumannii is a nonmotile, Gram-negative, nonfermentative, oxidase-negative, and aerobic Bacilli, which is one of the most opportunistic pathogens against human.[1,2] The bacteria are widespread in the environment and are considerably resistant to most antibiotics, low nutrient, and arid condition.[3] Acinetobacter spp. cause a variety of nosocomial infections, but A. baumannii is the prevalent species with high morbidity and mortality, including pneumonia, bacteremia, urinary tract, and skin and soft tissue infections, especially in patients with severe illness.[1,3,4] At the recent years, the continuously increasing prevalence rates of nosocomial infections by multidrug-resistant (MDR) A. baumannii in the Intensive Care Units (ICUs) have led to rise mortalities at the hospitals.[5-7]
A. baumannii has propensity to acquire resistance; a mechanism of this resistance is being characterized by the production of a specific enzyme called metallo-beta-lactamases (MβLs).[4,7] These enzymes belong to Ambler class B beta-lactamases based on their amino acid sequence homology and to Group 3 according to the Bush classification based on their substrate profiles (imipenem hydrolysis).[7] These enzymes are inhibited by ethylenediaminetetraacetic acid (EDTA).[8]
In Iran, several studies have previously been carried out on drug resistance of A. baumannii, revealing a high resistance rate to most of the antibiotics.[9-11] However, a few studies have been conducted in Iran, especially in our region, aiming to find the prevalence of MβL producing genes among clinical isolates of A. baumannii. The aims of the present study were to investigate MDR and determine the prevalence of MβL genes among carbapenem-resistant isolates of A. baumannii from clinical specimens in Shiraz, Southwest of Iran.
Materials and Methods
Sampling and bacterial isolates
In this cross-sectional study from October 2012 to April 2013, 120 Acinetobacter isolates were recovered from clinical specimens. The specimens were randomly collected from patients in different wards of two hospitals, Nemazee and Faghihi in Shiraz, a major city in Southwest of Iran. Nemazee and Faghihi hospitals are the two major tertiary care hospitals with 1000 beds, affiliated to Shiraz University of Medical Science, Shiraz, Iran. Sample size was calculated as about 100 using the formula [12]:
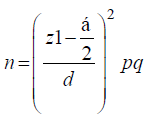
The study was in accordance with declaration of Helsinki; however, because we only used laboratory clinical specimens and did not harm any of the patients, the local ethics committee waived the need for informed consent.
Totally, 98 isolates were identified as A. baumannii isolates using Microgen™ diagnosis Kits. These 98 isolates were recovered from 35 sputum, 15 wound swab, 13 body fluids, 9 blood, 9 urine, 8 endotracheal tube, 5 cerebrospinal fluid, and 4 other samples (included 2 bronchoalveolar lavage, 1 eye swab, and 1 axillary swab). Overall, 53 samples were collected from ICUs and 45 samples from other wards (included gastroenterology, skin, surgery, and transplant) in the two above-mentioned hospitals. Confirmed isolates were kept at − 70°C until for long preservation.
Antimicrobial susceptibility testing
The susceptibility of the isolates to 16 antibiotics disk (MAST, UK) was investigated using disk diffusion method according to the Clinical and Laboratory Standards Institute (CLSI) recommendation.[13] The test was performed on Mueller-Hinton agar (Merck, Germany) with followed antibiotics disks containing 10 μg imipenem, 10 μg meropenem, 30 μg amikacin, 10 μg gentamicin, 30 μg aztreonam, 30 μg ceftazidime, 5 μg ciprofloxacin, 5 μg levofloxacin, 100 μg piperacillin, 10 μg tobramycin, 25 μg colistin, 20 μg ampicillin/sulbactam, 10 μg ampicillin, and 110 μg piperacillin/tazobactam. Pseudomonas aeruginosa ATCC 27853 was used as the control strain in susceptibility testing.
Minimal inhibitory concentration
Minimal inhibitory concentration (MIC)-test strips (Liofilchem, Italy), containing imipenem and meropenem, were used to determine the MIC according to the instruction provided by the company on Muller-Hinton agar (Merck, Germany). Results were interpreted using CLSI criteria.
Phenotypic metallo-beta-lactamase detection
MIC-test strip (Liofilchem, Italy), containing imipenem/ imipenem + EDTA, was used to determine the phenotypic MβL enzyme production. MIC-test strip was performed according to the manufacturer’s instructions. A reduction in the MIC of imipenem of ≥ 3 dilutions in the presence of EDTA was interpreted as a positive test. In addition, a strain was considered MβL producer if a phantom zone or deformation of the eclipse was observed.
DNA extraction and molecular typing
The isolates which were intermediately resistant or resistant to imipenem or meropenem were considered carbapenem-resistant. Extraction of genomic DNA from A. baumannii isolates was performed according to the protocol as previously described.[14] Polymerase chain reaction (PCR) assay was performed for the detection of blaOXA-51-like gene, a 353 base-pair (bp) amplicon, as an internal gene for molecular confirmation of A. baumannii isolates at the species level (primer sequence: F/5’-TAA TGC TTT GAT CGG CCT TG-3′, R/5′-TGG ATT GCA CTT CAT CTT GG-3′),[15] and for amplification of MβL encoding genes, blaIMP, blaVIM, and blaSPM using the following consensus primer sets, blaIMP: F/5’-GAA GGC GTT TAT GTT CAT AC-3’, blaIMP: R/5’-GTA TGT TTC AAG AGT GAT GC-3’ which amplify a 587 bp amplicon, blaVIM: F/5’-GTT TGG TCG CAT ATC GCA AC-3’, blaVIM: R/5’-AAT GCG CAG CAC CAG GAT AG-3’ which amplify a 382 bp amplicon,[16] and blaSPM: F/5’-AAA ATC TGG GTA CGC AAA CG-3’, blaSPM: R/5’-ACA TTA TCC GCT GGA ACA GG-3’ which amplify a 271 bp amplicon.[17] A P. aeruginosa harboring blaIMP, blaVIM, blaSPM genes (obtained from Pasteur Institute of Iran) was used as the positive control.[18]
Statistical analysis was performed using SPSS™ software version 19.0 (IBM Corp., Armonk, NY, USA). The results for infectious agents and antimicrobial susceptibility presented as descriptive statistics in terms of relative frequency. Chi-square or Fisher’s exact test was used to analyze the results wherever they needed. P < 0.05 was considered as statistically significant clinical relevance.
Results
Of 98 A. baumannii isolates, 54.1% (53/98) of isolates were collected from female patients’ samples. Seventy isolates were obtained from Namazee and 28 from Faghihi Hospital. Most of the 98 A. baumannii isolates were recovered from sputum (n = 35), wound swab (n = 15), and body fluids (n = 13).
The highest antibiotic resistance rates were observed against aztreonam, ceftazidime, ciprofloxacin, piperacillin, and cefotaxime since all of the tested isolates showed resistance. In addition, no sensitive isolates were seen against aztreonam and cefotaxime; just a few isolates were in the intermediate level of susceptibility. The results of antibiotic susceptibility patterns for the tested isolates are displayed [Table 1].
| Antibiotic disks | Total (n=98) n (%) | ||
|---|---|---|---|
| Sensitive | Intermediate | Resistance | |
| Meropenem | 3 (3.1) | 0 | 95 (96.9) |
| Imipenema | 4 (4.1) | 0 | 94 (95.9) |
| Amikacin | 24 (24.5) | 6 (6.1) | 68 (69.4) |
| Gentamicin | 13 (13.3) | 11 (11.2) | 74 (75.5) |
| Aztreonam | 0 | 0 | 98 (100) |
| Cefoxitin | 0 | 0 | 98 (100) |
| Ceftazidime | 0 | 0 | 98 (100) |
| Ciprofloxacin | 0 | 0 | 98 (100) |
| Levofloxacin | 0 | 0 | 98 (100) |
| Piperacillin | 0 | 0 | 98 (100) |
| Piperacillin-tazobactam | 0 | 0 | 98 (100) |
| Ampicillin | 0 | 0 | 98 (100) |
| Ampicillin-sulbactam | 5 (5.1) | 0 | 93 (94.9) |
| Colistin | 42 (42.9) | 0 | 56 (57.1) |
aResistance rate against imipenem using MIC was 99%, which is higher compared to 96% detected by disc diffusion method. MIC: Minimal inhibitory concentration.
Table 1: Susceptibility patterns of Acinetobacter baumannii isolates to 16 tested antibiotics by disc diffusion method.
Totally, 98% (96/98) of the isolates were detected as carbapenem-resistant by MIC-test strip. Among the carbapenem-resistant A. baumannii isolates, 44.8% (43/96) were found to be MβL producers by MβL MIC-test strips. Interestingly, the majority 85.7% (84/98) of the isolates showed an MIC ≥ 48 µg/ml. MIC ranges for the tested isolates to imipenem are presented in Table 2. All phenotypically MβL producing isolates were MDR and exhibited high resistance to beta-lactams, aminoglycosides, and fluoroquinolones. Furthermore, three of the MβL producing isolates were sensitive to ampicillin-sulbactam.
| MIC (μg/mL)sample | Sensitive≤4 μg/mL | Resistance≥16 μg/mL | ||||||
|---|---|---|---|---|---|---|---|---|
| 2 | 24 | 32 | 48 | 64 | 98 | 128 | Total number | |
| Sputum | 0 | 1 | 2 | 2 | 10 | 9 | 11 | 35 |
| Wound | 1 | 0 | 0 | 2 | 3 | 4 | 5 | 15 |
| Body fluids | 0 | 1 | 0 | 2 | 1 | 7 | 2 | 13 |
| Urine | 0 | 1 | 2 | 1 | 2 | 3 | 0 | 9 |
| Blood | 0 | 0 | 1 | 2 | 2 | 4 | 0 | 9 |
| ETTa | 0 | 2 | 1 | 2 | 2 | 1 | 0 | 8 |
| CSFb | 1 | 0 | 1 | 2 | 0 | 1 | 0 | 5 |
| Other | 0 | 0 | 0 | 2 | 1 | 0 | 1 | 4 |
| Total number | 2 | 5 | 7 | 15 | 21 | 29 | 19 | 98 |
aETT: Endotracheal tube, bCSF: Cerebrospinal fluid. MIC: Minimal inhibitory concentration.
Table 2: Distribution of resistance rate of the studied Acinetobacter baumannii isolates to imipenem by E-test based on source of isolation.
Finally from MβL producing isolates, 86% (37/43) were positive for MβL genotypes, of which 53.4% (23/43) isolates carried blaIMP, 32.6% (14/43) carried blaVIM MβL genes, and none of the isolates carried blaSPM gene. PCR results for amplification of MβL producing genes are displayed in Table 3.
| MβL gene | Sample | Place of isolation | Total number |
|---|---|---|---|
| IMP-1 | Sputum | ICU | 6 |
| IMP-1 | Sputum | Gastroenterology | 2 |
| IMP-1 | Body fluids | ICU | 2 |
| IMP-1 | Wound | ICU | 2 |
| IMP-1 | Wound | NICU | 1 |
| IMP-1 | Wound | Transplant | 1 |
| IMP-1 | Blood | ICU | 2 |
| IMP-1 | Blood | Transplant | 1 |
| IMP-1 | ETTa | NICU | 2 |
| IMP-1 | ETT | ICU | 1 |
| IMP-1 | Urine | ICU | 2 |
| IMP-1 | CSFb | ICU | 1 |
| VIM-2 | Sputum | ICU | 2 |
| VIM-2 | Sputum | Gastroenterology | 1 |
| VIM-2 | Wound | ICU | 2 |
| VIM-2 | Wound | NICU | 1 |
| VIM-2 | Urine | ICU | 2 |
| VIM-2 | Blood | ICU | 1 |
| VIM-2 | Blood | NICU | 1 |
| VIM-2 | Body fluids | ICU | 1 |
| VIM-2 | CSF | ICU | 1 |
| VIM-2 | ETT | Gastroenterology | 1 |
| VIM-2 | Other | NICU | 1 |
aETT: Endotracheal tube, bCSF: Cerebrospinal fluid. MßL: Metallo-beta-lactamase, ICU: Intensive Care Unit, NICU: Neonatal Intensive Care Unit
Table 3: Distribution of metallo-beta-lactamase genes among the studied Acinetobacter baumannii isolates
From the total ICUs recovered isolates, 47.2% (25/53) carried one of the blaIMP or blaVIM genes; this rate for isolates from other hospital wards was 26.7% (12/45). Despite the higher proportion for blaIMP and blaVIM gene in ICUs compared to other wards, no significant differences were observed (P = 0.06). The frequencies of blaIMP and blaVIM genes for Namazee hospital isolates were 22.9% (16/70) and 12.9% (9/70), respectively. The similar rates for Faghihi Hospital isolates were 25% (7/28) for blaIMP and 17.9% (5/28) for blaVIM genes.
Discussion
One of the carbapenem resistance mechanisms is production of carbapenem-hydrolyzing β-lactamases, which is called carbapenemase; one of them, MβL is more important in drug resistance against carbapenems.[7,19]
In the recent years, there have been numerous reports on MDR A. baumannii from hospital settings in Iran.[10-11] In most of these studies, only characterized isolates were obtained from ICUs while this study attempted to determine the resistance among isolates obtained from various medical wards as well. In particular, 45 clinical samples were isolated from wards outside ICUs. Antibiotic susceptibility testing showed that the majority of the isolates were resistant to three or more antibiotics [Table 1] while all the isolates were resistant to imipenem, cefotaxime, and ciprofloxacin, and 95% and 96% of the isolates were resistant to meropenem and amikacin, respectively.
Carbapenems are the best choice for nosocomial infections of Acinetobacter in Iran. However, in the recent years, it has been reported that there is reduced susceptibility to imipenem.[10,11,20]
Further, resistance to colistin, polymyxin, and tigecycline which are the usual choices for nosocomial infections of carbapenem-resistance Acinetobacter is increased around the world, especially in Iran.[10,11,21] Our results suggest colistin as the best choice for A. baumannii isolates in-vitro; this is in accordance with the finding of Japoni-Nejad et al. from Arak, Central area of Iran, that reported colistin and tigecycline as the most effective antibiotic agents against A. baumannii isolates.[10]
In a relatively similar study, Feizabadi et al. reported the prevalence of susceptibility of A. baumannii to imipenem, meropenem, piperacillin-tazobactam, and amikacin rate of 50.7%, 50%, 42.1%, and 38.2%, respectively.[22] These rates indicated higher sensitive isolates compared to our rates with 4.1%, 3.1%, 0%, and 24.5%, respectively. Our results showed a higher resistance rate of A. baumannii isolates against most of the studied antibiotics which were almost similar compared to some studies carried out throughout Iran and other countries.[23-25]
In this study, of the 96 (98%) carbapenem-resistant A. baumannii, 44.8% were found to be MβL producers by MIC-test strips. Such a high resistance rate to imipenem is not uncommon since two separate studies by Safari et al. and Noori et al. have previously shown 99% resistance among their tested isolated by MIC-test strips and disk diffusion methods, respectively.[11,21] Abdalhamid et al. from Saudi Arabia, in contrast with Iranian studies, reported lower carbapenem-resistant A. baumannii (46/141; 32.6%); however, MβL was much higher than that of Iranian report since 43 of 46 carbapenem-resistant A. baumannii isolates were MβL positive.[26]
All MβL producing isolates in our study were MDR and exhibited high resistance to beta-lactams, aminoglycosides, and fluoroquinolones. This high rate of MDR in our study was in agreement with a study from our neighboring country, Pakistan, which reported the prevalence of MDR 100% among A. baumannii isolates.[27] Three of our MβL-producing isolates were sensitive to ampicillin-sulbactam.
In the present study, from a total of 43 MβL producing isolates, 86% were positive for MβL genotypes and subsequently 53.4% and 32.6% of the isolates carried blaIMP and blaVIM genes, respectively.
As it is seen in our results, phenotypic and genotypic results of the detection of MβL producing isolates were not the same. Possible reason of higher positive rates for phenotypic MβL production could be the presence of MβL genes other than those we have screened.[28,29]
In a previous Iranian study conducted in Tabriz, from a total of 63 carbapenem-resistant A. baumannii, 31 (49%) were found to be MβL producers by MIC-test strips, of which 19 isolates carried blaIMP and 9 carried blaVIM genes; these rates were remarkably close to our findings.[20] In addition, Noori et al. from the capital of Iran reported a blaIMP-1 gene prevalence of 3 of 86 (3.48%) among MβL A. baumannii isolates.[21]
In the current study, no blaSPM gene was detected among A. baumannii isolates; previously, two similar attempts for detection of blaSPM gene by Shahcheraghi et al. (2009–2010) and Noori et al. (2012–2013) were made in Tehran, Iran; only, Shahcheraghi et al. was able to detect this gene in 6 out of 100 tested A. baumannii isolates.[18,21] Reports about the prevalence of these genes among A. baumannii isolates from other parts of the world have shown different patterns. Abdalhamid et al. from Saudi Arabia and Al-Agamy et al. from Egypt reported negative PCR results for blaIMP, blaVIM, and blaSPM genes among all MβL producing isolates.[26,30] However, previously, blaIMP and blaVIM prevalences have been reported from Greece, China, Korea, and India.[31-34] The present study has some limitations. First, lack of DNA sequencing for knowing the IMP and VIM variants must be mentioned as a limitation of our study. Moreover, employed a typing method could clear the connection between isolates and source of infections in studied hospitals.
Conclusion
The prevalence of drug resistance among the clinical samples of A. baumannii isolated in our study against most of the antibiotics is very high. Hence, early detection and optimization of infection control practices are the best defenses against these organisms. Therefore, systematic surveillance to detect MβL producing bacteria and rational prescription and use of carbapenems could be helpful to prevent the spread of carbapenem resistance.
Acknowledgements
The authors would like to thank Ms. N. Pirbonyeh and Mr. B. Dehghani for their technical assistance. This study was supported by Shiraz University of Medical Sciences with grant No. 92-6515. This article is extracted from the MSc thesis by M. N. Moghadam under the supervision of Dr. M. Motamedifar.
Financial support and sponsorship
This study was supported by Shiraz University of Medical Sciences with grant No. 92-6515.
Conflicts of interest
There are no conflicts of interest.
References
- Karageorgopoulos DE, Kelesidis T, Kelesidis I, Falagas ME.Tigecycline for the treatment of multidrug-resistant (including carbapenem-resistant) Acinetobacter infections: A review of the scientific evidence. J Antimicrob Chemother 2008;62:45-55.
- Karah N, Sundsfjord A, Towner K, Samuelsen Ø. Insights into the global molecular epidemiology of carbapenem non-susceptible clones of Acinetobacter baumannii. Drug Resist Updat 2012;15:237-47.
- Cisneros JM, Reyes MJ, Pachón J, Becerril B, Caballero FJ,García-Garmendía JL, et al. Bacteremia due to Acinetobacter baumannii: Epidemiology, clinical findings, and prognostic features. Clin Infect Dis 1996;22:1026-32.
- Kuo SC, Chang SC, Wang HY, Lai JF, Chen PC, Shiau YR, et al. Emergence of extensively drug-resistant Acinetobacter baumannii complex over 10 years: Nationwide data from
- the Taiwan surveillance of antimicrobial resistance (TSAR) program. BMC Infect Dis 2012;12:200.
- Kooti S, Motamedifar M, Sarvari J. Antibiotic resistance profile and distribution of oxacillinase genes among clinical isolates of Acinetobacter baumannii in Shiraz teaching hospitals,2012-2013. Jundishapur J Microbiol 2015;8:e20215.
- Zheng W, Yuan S, Li L. Analysis of hospital departmental distribution and antibiotic susceptibility of Acinetobacter isolated from sputum samples. Am J Infect Control 2013;41:e73-6.
- Maltezou HC. Metallo-beta-lactamases in Gram-negative bacteria: Introducing the era of pan-resistance? Int J Antimicrob Agents 2009;33:405.e1-7.
- Yong D, Lee K, Yum JH, Shin HB, Rossolini GM, Chong Y.Imipenem-EDTA disk method for differentiation of metallo-beta-lactamase-producing clinical isolates of Pseudomonas spp. and Acinetobacter spp. J Clin Microbiol 2002;40:3798-801.
- HakemiVala M, Hallajzadeh M, Fallah F, Hashemi A, Goudarzi H. Characterization of the extended-spectrum beta-lactamase producers among non-fermenting Gram-negative bacteria isolated from burnt patients. Arch Hyg Sci 2013;2:1-6. 10. Japoni-Nejad A, Sofian M, Belkum AV, Ghaznavi-Rad E. Nosocomial outbreak of extensively and pan drug-resistant Acinetobacter baumannii in tertiary hospital in central part of Iran. Jundishapur J Microbiol. 2013;6:e9892.
- Safari M, Saidijam M, Bahador A, Jafari R, Alikhani MY. High prevalence of mul t idrug resistance and metallo-beta-lactamase (MßL) producing Acinetobacter baumannii isolated from patients in ICU wards, Hamadan, Iran. J Res Health Sci 2013;13:162-7.
- Oladeinde B, Ekejindu I, Omoregie R, Aguh O. Awareness and knowledge of ergonomics among Medical Laboratory Scientists in Nigeria. Ann Med Health Sci Res. 2015;5:423-7.
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing; 24th Informational Supplement. M100-S24. Wayne, PA: Clinical and Laboratory Standards Institute; 2014.
- Sambrook J, Russell DW. Purification of nucleic acids by extraction with phenol: Chloroform. CSH Protoc 2006;2006. pii: pdb.prot4455.
- Turton JF, Woodford N, Glover J, Yarde S, Kaufmann ME, Pitt TL. Identification of Acinetobacter baumannii by detection of the blaOXA-51-like carbapenemase gene intrinsic to this species. J Clin Microbiol 2006;44:2974-6.
- Khosravi AD, Mihani F. Detection of metallo-beta-lactamaseproducing Pseudomonas aeruginosa strains isolated from burn patients in Ahwaz, Iran. Diagn Microbiol Infect Dis 2008;60:125-8.
- Poirel L, Walsh TR, Cuvillier V, Nordmann P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn Microbiol Infect Dis 2011;70:119-23.
- Shahcheraghi F, Abbasalipour M, Feizabadi M, Ebrahimipour G, Akbari N. Isolation and genetic characterization of metallo-ß-lactamase and carbapenamase producing strains of Acinetobacter baumannii from patients at Tehran hospitals. Iran J Microbiol 2011;3:68-74.
- Ben RJ, Yang MC, Hsueh JC, Shiang JC, Chien ST. Molecular characterisation of multiple drug-resistant Acinetobacter baumannii isolates in southern Taiwan. Int J Antimicrob Agents 2011;38:403-8.
- Peymani A, Nahaei MR, Farajnia S, Hasani A, Mirsalehian A, Sohrabi N, et al. High prevalence of metallo-beta-lactamase-producing Acinetobacter baumannii in a teaching hospital in Tabriz, Iran. Jpn J Infect Dis 2011;64:69-71.
- Noori M, Karimi A, Fallah F, Hashemi A, Alimehr S, Goudarzi H, et al. High prevalence of metallo-beta-lactamase producing Acinetobacter baumannii isolated from two hospitals of Tehran, Iran. Arch Pediatr Infect Dis 2014;2:e15439.
- Feizabadi MM, Fathollahzadeh B, Taherikalani M, Rasoolinejad M, Sadeghifard N, Aligholi M, et al. Antimicrobial susceptibility patterns and distribution of blaOXA genes among Acinetobacter spp. Isolated from patients at Tehran hospitals. Jpn J Infect Dis 2008;61:274-8.
- Rahbar M, Mehrgan H, Aliakbari NH. Prevalence of antibiotic-resistant Acinetobacter baumannii in a 1000-bed tertiary care hospital in Tehran, Iran. Indian J Pathol Microbiol 2010;53:290-3.
- Akbari M, Niakan M, Taherikalani M, Feizabadi MM, Azadi NA, Soroush S, et al. Rapid identification of Iranian Acinetobacter baumannii strains by single PCR assay using BLA oxa-51-like carbapenemase and evaluation of the antimicrobial resistance profiles of the isolates. Acta Microbiol
- Immunol Hung 2010;57:87-94. 25. Hamzeh AR, Al Najjar M, Mahfoud M. Prevalence of antibiotic resistance among Acinetobacter baumannii isolates from Aleppo, Syria. Am J Infect Control 2012;40:776-7.
- Abdalhamid B, Hassan H, Itbaileh A, Shorman M.Characterization of carbapenem-resistant Acinetobacter baumannii clinical isolates in a tertiary care hospital in Saudi Arabia. New Microbiol 2014;37:65-73.
- Begum S, Hasan F, Hussain S, Ali Shah A. Prevalence of multi drug resistant Acinetobacter baumannii in the clinical samples from tertiary care hospital in Islamabad, Pakistan. Pak J Med Sci 2013;29:1253-8.
- Meletis G, Exindari M, Vavatsi N, Sofianou D, Diza E.Mechanisms responsible for the emergence of carbapenem resistance in Pseudomonas aeruginosa. Hippokratia 2012;16:303-7.
- Poirel L, Nordmann P. Carbapenem resistance in Acinetobacter baumannii: Mechanisms and epidemiology. Clin Microbiol Infect 2006;12:826-36.
- Al-Agamy MH, Khalaf NG, Tawfick MM, Shibl AM, ElKholy A. Molecular characterization of carbapenem-insensitive Acinetobacter baumannii in Egypt. Int J Infect Dis 2014;22:49-54.
- Tsakris A, Ikonomidis A, Pournaras S, Tzouvelekis LS, Sofianou D, Legakis NJ, et al. VIM-1 metallo-beta-lactamase in Acinetobacter baumannii. Emerg Infect Dis 2006;12:981-3.
- Chen Z, Qlu S, Wang Y, Wang Y, Liu S, Wang Z, et al. Coexistence of blaNDM-1 with the prevalent blaOXA23 and blaIMP in pan-drug resistant Acinetobacter baumannii isolates in China. Clin Infect Dis 2011;52:692-3.
- Lee K, Lee WG, Uh Y, Ha GY, Cho J, Chong Y; Korean Nationwide Surveillance of Antimicrobial Resistance Group. VIM- and IMP-type metallo-beta-lactamase-producing Pseudomonas spp. and Acinetobacter spp. in Korean hospitals. Emerg Infect Dis 2003;9:868-71.
- Amudhan MS, Sekar U, Kamalanathan A, Balaraman S.bla (IMP) and bla (VIM) mediated carbapenem resistance in Pseudomonas and Acinetobacter species in India. J Infect Dev Ctries 2012;6:757-62.



 The Annals of Medical and Health Sciences Research is a monthly multidisciplinary medical journal.
The Annals of Medical and Health Sciences Research is a monthly multidisciplinary medical journal.